How-to simulate Support Vector Machine (SVM) in R Science 22.04.2014
 Introduction
Introduction
Support Vector Machine (SVM) is a supervised machine learning algorithm which can be used for classification or regression problems (numeric prediction).
Given a set of training examples, where each data point falls into one of two categories, an SVM training algorithm builds a model that assigns new data points into one category or the other. This model is a representation of the examples as a points in space, mapped so that the examples of the separate categories are divided by a margin that is as wide as possible, as shown in the following image. New examples are then mapped into that same space and predicted to belong to a category based on which side of the gap they fall on.
The approach creates a hyperplane to categorize the training data. A hyperplane can be envisioned as a geometric plane that separates two regions. In a two-dimensional space, it will be a line. In a three-dimensional space, it will be a two-dimensional plane. For higher dimensions, it is harder to conceptualize, but they do exist.
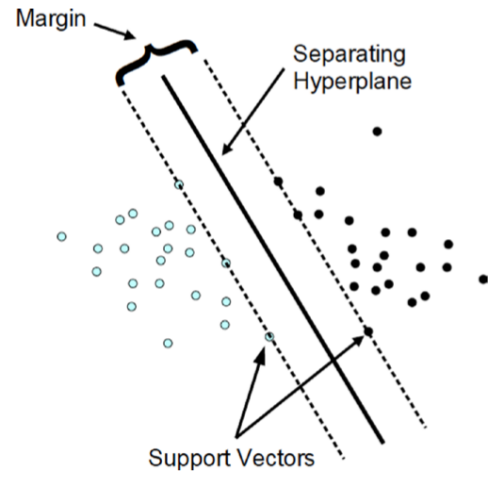
SVM looks for the optimal hyperplane separating the two classes by maximizing the margin between the closest points of the two classes. In two-dimensional space, as shown in figure below, the points lying on the margins are called support vectors and line passing through the midpoint of margins is the optimal hyperplane. In two dimensions, a hyperplane is a line and in a p-dimensional space, a hyperplane is a flat affine subspace of hyperplane dimension p - 1.
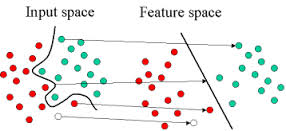
A SVM can be imagined as a surface that creates a boundary between points of data plotted in multidimensional that represent examples and their feature values. The goal of a SVM is to create a flat boundary called a hyperplane, which divides the space to create fairly homogeneous partitions on either side. In this way, the SVM learning combines aspects of both the instance-based nearest neighbor learning and the linear regression modeling. The combination is extremely powerful, allowing SVMs to model highly complex relationships.
Support vectors are data points that lie near the hyperplane. SVM uses a technique called the kernel trick to transform your data and then based on these transformations it finds an optimal boundary between the possible outputs. Simply put, it does some extremely complex data transformations, then figures out how to seperate your data based on the labels or outputs you've defined.
A few of the most commonly used kernel functions are listed as follows.
- The linear kernel does not transform the data at all.
- The polynomial kernel of degree d adds a simple nonlinear transformation of the data.
- The sigmoid kernel results in a SVM model somewhat analogous to a neural network using a sigmoid activation function.
- The Gaussian RBF kernel is similar to a RBF neural network. The RBF kernel performs well on many types of data and is thought to be a reasonable starting point for many learning tasks.
Math background you can find here.
Advantages and Disadvantages of SVM
An important benefit of the support vector machine approach is that the complexity of the resulting classifier is characterised by the number of support vectors rather than the dimensionality of the transformed space. As a result, support vector machines tend to be less prone to problems of over-fitting than some other methods.
There are four main advantages: Firstly it has a regularisation parameter, which makes the user think about avoiding over-fitting. Secondly it uses the kernel trick, so you can build in expert knowledge about the problem via engineering the kernel. Thirdly an SVM is defined by a convex optimisation problem (no local minima) for which there are efficient methods (e.g. SMO). Lastly, it is an approximation to a bound on the test error rate, and there is a substantial body of theory behind it which suggests it should be a good idea. Source.
The disadvantages are that the theory only really covers the determination of the parameters for a given value of the regularisation and kernel parameters and choice of kernel. In a way the SVM moves the problem of over-fitting from optimising the parameters to model selection. Sadly kernel models can be quite sensitive to over-fitting the model selection criterion, see
Compare SVM with NN (neural network) (source)
Similarity
- SVM + sigmoid kernel ~ two-layer feedforward NN
- SVM + Gaussian kernel ~ RBF network
- For most problems, SVM and NN have similar performance
Advantages
- Based on sound mathematics theory
- Learning result is more robust
- Over-fitting is not common
- Not trapped in local minima (because of QP)
- Fewer parameters to consider (kernel, error cost C)
- Works well with fewer training samples (number of support vectors do not matter much)
- Can be used for classification or numeric prediction problems
- May be easier to use than neural networks, particularly due to the existence of several well-supported SVM algorithms
Disadvantages
- Problem need to be formulated as 2-class classification
- Learning takes long time (QP optimization)
- Difficult to understand the learned function (weights)
- Finding the best model requires testing of various combinations of kernels and model parameters
- Can be slow to train, particularly if the input dataset has a large number of features or examples
- Results in a complex black box model that is difficult, if not impossible, to interpret
Multi-Class SVM
Most popular usage of SVM is as a binary classifier. However the idea of SVM can be extended to multi-class classification problem as well. Multi-class SVM can be used as a multi-class classifier by creating multiple binary classifiers. This method works similarly to the idea of multinomial logistic regression, where we build a logistic model for each pair of class with base function.
We can create a set of binary SVMs to do multi-class classification. The steps in implementing that will be as follows:
- Create binary classifiers:
- Between one class and the rest of the classes.
- Between every pair of classes (all possible pairs).
- For any new cases, the SVM classifier adopts a winner-takes-all strategy, in which the class with highest output is assigned.
To implement this methodology, it is important that the output functions are calibrated to generate comparable scores, otherwise the classification will become biased.
There are other methods also for multi-class SVMs. One such is proposed by Crammer and Singer. They proposed a multi-class SVM method which casts the multiclass classification problem into a single optimization problem, rather than decomposing it into multiple binary classification problems.
We will show quick example with iris data. The iris$Species is divided into three classes: setosa, versicolor, and virginica. The multi-class SVM has to classify iris into these categories.
Here is the R implementation of the SVM multi-class classifier:
#install.packages("e1071")
library("e1071")
iris_mdl = svm(Species ~ ., data=iris, header=T)
iris_predict = predict(iris_mdl, newdata=iris)
table(iris$Species, iris_predict)
classification_rate = sum(diag(table(iris$Species, iris_predict))) / nrow(iris)
Multi-class SVM gives us 97% good classification_rate on iris data. The prediction is good across all the classes.
A number of applications of SVM emerged over the years, and a few noteworthy ones are hypertext categorization, image classification, character recognition, and many more applications in biological sciences as well.
Modeling SVM in R
Good explanation of modeling SVM in R you can find in
- Alexandros Karatzoglou, David Meyer, Kurt Hornik. Support Vector Machines in R
- Jean-Philippe Vert. Practical session: Introduction to SVM in R
- Basic Classification in R
- Predictive Analytics: NeuralNet, Bayesian, SVM, KNN
Useful links
Quote
Categories
- Android
- AngularJS
- Databases
- Development
- Django
- iOS
- Java
- JavaScript
- LaTex
- Linux
- Meteor JS
- Python
- Science
Archive ↓
- September 2024
- December 2023
- November 2023
- October 2023
- March 2022
- February 2022
- January 2022
- July 2021
- June 2021
- May 2021
- April 2021
- August 2020
- July 2020
- May 2020
- April 2020
- March 2020
- February 2020
- January 2020
- December 2019
- November 2019
- October 2019
- September 2019
- August 2019
- July 2019
- February 2019
- January 2019
- December 2018
- November 2018
- August 2018
- July 2018
- June 2018
- May 2018
- April 2018
- March 2018
- February 2018
- January 2018
- December 2017
- November 2017
- October 2017
- September 2017
- August 2017
- July 2017
- June 2017
- May 2017
- April 2017
- March 2017
- February 2017
- January 2017
- December 2016
- November 2016
- October 2016
- September 2016
- August 2016
- July 2016
- June 2016
- May 2016
- April 2016
- March 2016
- February 2016
- January 2016
- December 2015
- November 2015
- October 2015
- September 2015
- August 2015
- July 2015
- June 2015
- February 2015
- January 2015
- December 2014
- November 2014
- October 2014
- September 2014
- August 2014
- July 2014
- June 2014
- May 2014
- April 2014
- March 2014
- February 2014
- January 2014
- December 2013
- November 2013
- October 2013